RNPxl has been integrated into OpenMS and also the Proteome Discoverer Community Nodes. No custom installer is required anymore.
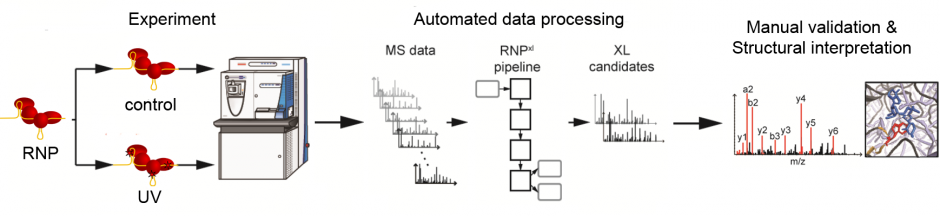
RNPxl has been integrated into OpenMS and also the Proteome Discoverer Community Nodes. No custom installer is required anymore.
Requirements:
- HCD data acquired on a high-resolution MS
- Developed and tested on orbitrap instruments
- MS and MS/MS need to be centroided (either on acquisition, conversion or in a workflow using the TOPP tool HiResPeakPicker)
For additional information on sample handling and acquisition, as well as a detailed step-by-step tutorial, please refer to the supplementary material of the publication.
Raw files:
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the PRIDE partner repository with the dataset identifier PXD000513.
PUBLICATION:
[Bibtex]
@article{kramer2014photo,
title={Photo-cross-linking and high-resolution mass spectrometry
for assignment of RNA-binding sites in RNA-binding proteins},
author={Kramer, Katharina and Sachsenberg, Timo and Beckmann, Benedikt M and Qamar, Saadia and Boon,
Kum-Loong and Hentze, Matthias W and Kohlbacher, Oliver and Urlaub, Henning},
journal={Nature methods},
volume={11},
number={10},
pages={1064--1070},
year={2014},
publisher={Nature Publishing Group}
}
FAQ:
- Q: RNPxlcrashes with error message indicating that an idXML was not found:
Error: File not found (the file 'C:/Users/username/AppData/Local/Temp/MSfilename_Number_Number_variant.idXML' could not be found)
RNPxl failed!
-
A: OMSSA has problems reading the path to the database. Check that the path to your database or windows temp directory doesn’t contain any spaces or additional dots. Make sure that all database related files (.fasta, .phr, .pin, .psq) are in the same folder and have the same name that you defined both in the parameters of the RNPxl tool and the OMSSA .ini file. Also make sure there is no dot except for the one between file name and file type, e.g., do not call the database “uniprot.human.fasta”
-
Q: Can you provide some details on the mass and losses of 4-thio-U nucleotide?
-
A: The monoisotopic mass of a 4-thio-U nucleotide is: 340.01301 (C9H13N2O8PS).However, a net loss of H2S has always been observed in our cross-linking experiments, making the adduct mass: 306.02529 (C9H11N2O8P). This adduct is observed on precursor level. Fragmentation cleaves off phosphate and ribose, leaving 94.01671 (C4H2N2O) as adduct on MS2 level.
If you are interested in the original data, binaries and workflows you may download:
Download original sample data and TOPPAS workflows:
RNPxl samples and pipelines Please note that RNPxl_sample_UV.mzML and RNPxl_sample_control.mzML only contain a portion of a full experiment and have already been picked to reduce the size of the file.